Saarbrücken, 8 December 2021 – an active compound can only exert its desired effects, if it reaches its target site. While this statement seems straightforward, the science behind it can be challenging. Understanding how active compounds permeate across the bacterial cell envelope and having tools to properly assess this process is essential – especially when developing novel antibiotics. Scientists from the Department of Drug Delivery across Biological Barriers and the Department of Microbial Natural Products at HIPS have now developed an Outer Membrane Vesicles (OMVs) Permeation Assay, which could serve as a high-throughput in vitro screening tool to assess compound- and bacteria-specific passive uptake pathways. They report about this assay in the journal Advanced Healthcare Materials.
Your research article is about an Assay for assessing bacterial bioavailability. Why is it so important to have a tool to determine an antibiotics permeability and what makes your assay special in comparison to previous approaches?
Estimating the uptake of antibiotic candidates is often crucial for their further development, since sufficient availability at the target site - one could speak of ‘bacterial bioavailability’ - determines their activity. Especially, bacteria with a double cell membrane (so called ‘Gram-negative bacteria’) can effectively control antibiotic concentration inside the cell to avoid reaching a harmful level. To this end, they use for example membrane-associated porin proteins, lipopolysaccharides and efflux pumps. Whole cell-based assays are quite bacteria-specific, require high-end analytical devices and commonly bear an infection hazard. Furthermore, it is difficult to rule out certain accumulation-determining factors. This makes these assays very complex and not suitable as a high-throughput screening method. Other cell-free assays are highly simplified in their composition, fragile, and also not suitable for high-throughput. Our OMV-based model gives the opportunity to estimate the permeability of antibiotics across the outer membrane, which is a very important factor contributing to bacterial bioavailability. It potentially leads to authentic permeation data with a minimum demand on work force and analytical equipment. This makes it applicable in many different labs.
Can you tell us a bit more about extracellular vesicles and what makes them interesting for your research?
Extracellular vesicles are like tiny liquid-filled bubbles. They are surrounded by a biomembrane and are released by virtually all types of cells to their environment. They fulfil various tasks, for example horizontal gene transfer, cell-to-cell communication, defense against predatory microorganisms or immune modulation. OMVs are one type of extracellular vesicles. They originate from the outer membrane of Gram-negative bacteria and feature typical membrane components like phospholipids, lipopolysaccharides, porins as well as fragments of active uptake or efflux transporters. This is what makes them attractive as a material for an in vitro membrane permeation model.
How did you characterize these OMVs as well as the in vitro assay?
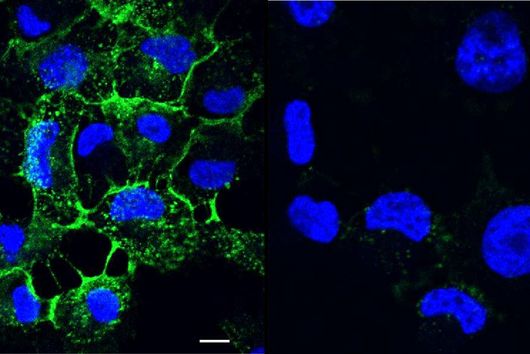
Due to the complexity of OMVs, we needed various characterization methods. The workflow started with dynamic light scattering and was followed by laser-doppler-anemometry, nanoparticle tracking analysis to characterize their size, surface charge and concentration. We then used microscopy-based methods, such as confocal laser scanning microscopy, scanning electron microscopy, cryo-transmission electron microscopy to characterize the shape and merging behaviour. In addition, protein analytics like SDS-PAGE, western blot and MALDI-TOF-Mass spectrometry were applied to confirm that the material we were dealing with was indeed consisting of outer membrane vesicles.
Preparing an OMV-based membrane model seems to be quite an intensive process; could you take us through the workflow?
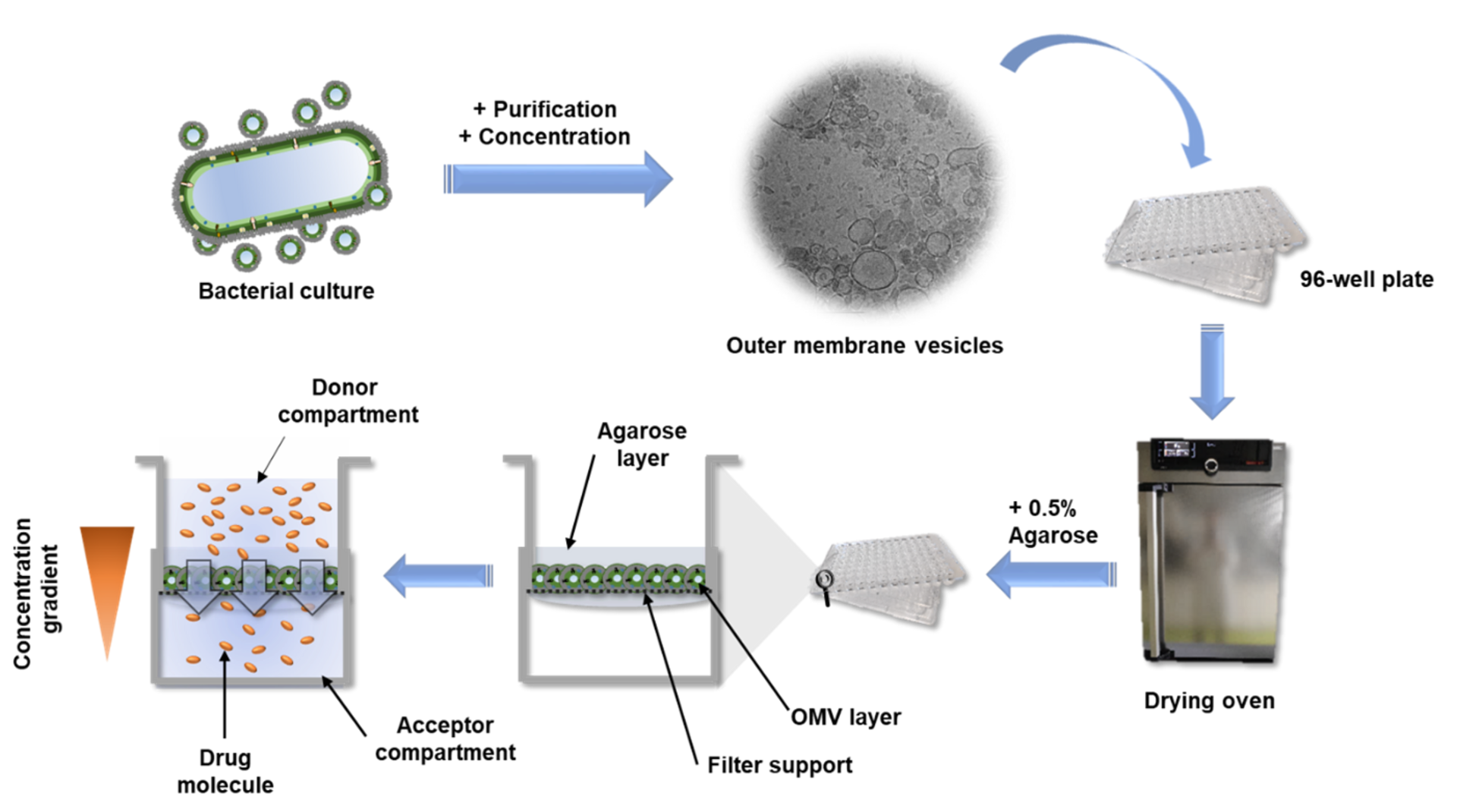
Compared to the tedious characterizations of this material, the preparation of the OMV-based model is rather simple: a bacterium of interest is kept in liquid culture over a defined amount of time. Afterwards, the OMV-containing medium is separated from the bacteria by centrifugation and sterile filtration. To separate the OMV from the medium, we decided to precipitate the OMVs using a concentrated PEG (polyethylene glycol) solution. PEG breaks the hydrogen bonds between OMVs and water, so that a normal centrifugation step is sufficient to isolate the OMV pellet from the liquid. After reconstituting the vesicles in buffer, they are dispensed multiple times on filter supports and dried under airflow. To enhance the model stability, a low-concentrated agarose gel is given on top, et voilà: the OMV-based membrane model is ready.
In your article, you mention that this assay could serve as a high-throughput screening tool. Can you explain how this would work and how other scientists could benefit from your assay?
The high-throughput of this assay is possible due to the option to quickly read out the compound absorbance or fluorescence directly from the 96-well receiver plate, on which the filter supports are located. Alternatively, the receiver compartments can be converted into donor compartments and sampling can be done in situ from the top of the coated filter supports via liquid handling systems. Such samples can then either be quantified by UV-vis spectrometry, fluorimetry or chromatographic methods. The dependency of this assay on very basic materials allows it to be implemented in various labs and the application is easy to learn. Hence, other scientists can easily test their new drug candidates on their self-made OMV-based model and maybe even further optimize and characterize it. The ‘Linux’ among the in vitro assays one could say.
Dr Alwin Hartman and Dr Yannic Nonnenmacher conducted the interview.
ORIGINAL PUBLICATION
Richter, R., Kamal, M.A.M., Koch, M. et al. An Outer Membrane Vesicle-Based Permeation Assay (OMPA) for Assessing Bacterial Bioavailability. Advanced Healthcare Materials (2021). DOI: 10.1002/adhm.202101180