Saarbrücken, January 09, 2024 - Modern pharmaceutical research generates large amounts of data faster than ever before. To analyze and evaluate this data, it is collected and structured in extensive databases and thus provides the basis for the development of new active ingredients. Researchers at the Helmholtz Institute for Pharmaceutical Research Saarland (HIPS) have now published two such databases. While the "ABC-HuMi" database focuses on genetic data from microorganisms that colonize humans, "ZEBRA" provides a high-resolution cell atlas of the human and mouse brain. Both databases were published in the annual database issue of the journal Nucleic Acids Research.
Scientific experiments generally not only lead to new findings but also uncover new questions and unsolved problems. To be able to process these efficiently, both further experiments as well as computer-based tools are required. In addition to software tools, this also includes databases that present research data in a structured way and make it accessible to the scientific community. The two HIPS scientists Prof Alexey Gurevich, head of the Human-Microbe Systems Bioinformatics group, and Dr Fabian Kern, head of the junior research group Spatiotemporal Single-Cell Bioinformatics, have successfully published two of their databases in the annual database issue of the renowned journal Nucleic Acids Research. This special issue is one of the most important outlets for the publication and dissemination of bioinformatics databases. The HIPS is a site of the Helmholtz Centre for Infection Research in collaboration with Saarland University.
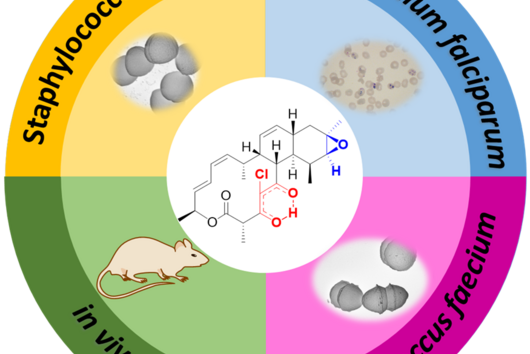
The ABC-HuMi database developed by Gurevich and colleagues contains an extensive collection of genetic data on human microbiota, i.e. microorganisms that colonize the human body. A special focus of ABC-HuMi is on so-called biosynthetic gene clusters. These genetic sections contain blueprints for the production of natural products with which microorganisms communicate or fight each other. Such substances also play a major role in the interaction with the human host, particularly during the emergence and development of various diseases. "Our overarching goal is to understand the interaction between humans and their microbiota on a chemical level. To do this, we first need to know which chemical signals are involved in the interaction and elucidate their roles. We then set out to use this information to develop new active substances. ABC-HuMi lays the foundation for this endeavor," says Alexey Gurevich. A special feature of the database is that it combines data from microbiota from different areas of the human body and does not just focus on areas that are already well characterized, such as the intestine.
In a second database, entitled ZEBRA, a team led by HIPS junior research group leader Fabian Kern analyzed the human and mouse brain down to the level of individual cells. ZEBRA allows investigating the activity of different genes in specific areas of the brain or even single cells - depending on factors such as age, sex or health status. In total, the database contains information on 4.2 million cells from 39 brain regions. These detailed insights into the activity of the most complex human organ should help to better understand the largely unknown causes and mechanisms behind neurodegenerative diseases. Infection research can also benefit from the new database: "In the case of SARS-CoV-2, it was initially assumed that the virus mainly damages the respiratory tract. However, in one of our previous studies, we were able to show that Covid-19 infection also induces inflammation in the brain. ZEBRA includes data from such patients and thus enables us to find out what exactly happens in the brain during infection and to develop strategies to counteract these effects," says Fabian Kern.
Andreas Keller, Professor at Saarland University and head of the Clinical Bioinformatics department at HIPS, emphasizes the importance of the two publications for a bioinformatics-backed strategy at the Saarbrücken site: "Our aim is to establish an ecosystem in the field of bioinformatics in Saarbrücken that is closely networked with experimental research. The published databases are an important step in this direction and also serve as reference data sets for scientists all over the world. The databases and software applications we develop enable increased interaction between different research disciplines. It is very important to us that all resources are openly accessible to researchers all around the world." The field of bioinformatics will be further expanded at HIPS and will play a key role at the institute. The Scientific Director of HIPS, Prof Rolf Müller, says: "Over the last two years, we have succeeded in building up a young and excellent team in the field of bioinformatics under the leadership of Andreas Keller, which covers a broad spectrum of expertise. The integration of bioinformaticians into our research projects is crucial to unlocking the full potential of our data. Especially at the interface between drug discovery and medicine, we see an opportunity to significantly advance our research through the implementation of computer science."
The databases can be accessed via the following links:
ABC-HuMi: https://www.ccb.uni-saarland.de/abc_humi/
ZEBRA: https://www.ccb.uni-saarland.de/zebra
Original publications:
Pascal Hirsch, Azat Tagirdzhanov, Aleksandra Kushnareva, Ilia Olkhovskii, Simon Graf, Georges P Schmartz, Julian D Hegemann, Kenan A J Bozhüyük, Rolf Müller, Andreas Keller, Alexey Gurevich, ABC-HuMi: the Atlas of Biosynthetic Gene Clusters in the Human Microbiome, Nucleic Acids Research, 2023; DOI: 10.1093/nar/gkad1086
Matthias Flotho, Jérémy Amand, Pascal Hirsch, Friederike Grandke, Tony Wyss-Coray, Andreas Keller, Fabian Kern, ZEBRA: a hierarchically integrated gene expression atlas of the murine and human brain at single-cell resolution, Nucleic Acids Research, 2023; DOI: 10.1093/nar/gkad990